MENU
US | USD
US | USD
-
- All Centrifuges
- Benchtop Centrifuges
- Floor-Standing Centrifuges
- Refrigerated Centrifuges
- Microcentrifuges
- Multipurpose Centrifuges
- High-Speed Centrifuges
- Ultracentrifuges
- Concentrator
- High-Speed and Ultracentrifuge Consumables
- Centrifuge Tubes
- Centrifuge Plates
- Device Management Software
- Sample and Information Management
-
- All Pipettes, Dispensers & Automated Liquid Handlers
- Mechanical Pipettes
- Electronic Pipettes
- Multi-Channel Pipettes
- Positive Displacement Pipettes & Dispensers
- Pipette Tips
- Bottle-Top Dispensers
- Pipette Controllers
- Dispenser & Pipette Accessories
- Automated Pipetting
- Automation Consumables
- Automation Accessories
- Liquid Handler & Pipette Services
Sorry, we couldn't find anything on our website containing your search term.
Sorry, we couldn't find anything on our website containing your search term.
Gradient PCR - Optimizing temperatures and elongation time in PCR
Lab Academy
- Molecular Biology
- Amplification & PCR
- PCR Cyclers
- Essay
Anyone who has performed PCR would be familiar with the three basic steps in the “chain reaction” – denaturation, annealing and extension (also known as elongation). In general, the temperature used in the denaturation step is dependent on the DNA.
The extension temperature is generally set at 72°C because Taq polymerase works optimally at this temperature. The duration of the extension step also matters, and is typically about one minute per kilobase of target length. However, modern high-speed enzymes can shorten the elongation time in PCR significantly.
Read more
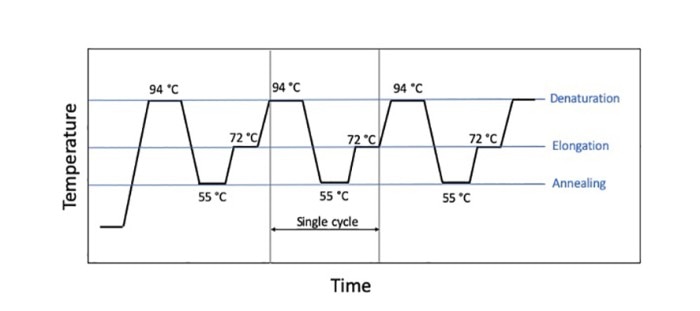
Figure 1: The three main steps of a PCR style, denaturation, extension, and annealing, with their standard temperatures given.
This means at each PCR step, there is a degree of flexibility. The positive consequence of this is versatility – temperatures and durations can be modified to suit almost any applications, steps can be combined to shorten the total reaction time, multiple PCRs can be run together using a shared condition, etc. However, the problem with any versatility is, finding the “best” condition becomes almost an art. If everything is possible, finding “the one” is like searching for a single fish in an ocean. Fortunately, there are some guidelines to follow, meaning it is not to the point of despair!
For example, we know that the denaturation temperature should be as low as reasonable to denature template DNA and thus allowing efficient primer annealing and strand extension1 . At the same time, it should not be too high to prevent loss of enzyme activity, since the half-life of Taq DNA polymerases drastically reduces with increasing temperature2 . Naturally, decades of improvements have resulted in more and more stable PCR reagent formulations – increasing fidelity, specificity and speed performance of enzymes. Thus, the denaturation temperature generally falls between 92°C to 98°C, offering choices based on users’ applications. For example, difficult templates such as GC-rich sequences require a higher denaturation temperature, which not only helps separate the strands, but can also lead to higher specificity in PCR amplification3 .
Things become more complicated when choosing the right annealing temperature. The closest reference one has is the melting temperatures (TM) of the forward and reverse primers. TM describe the temperature at which there is an equilibrium between the primer bound to DNA and the primer dissociating from DNA, resulting in 50% of the template having bound primer. When choosing primer pairs, it is important that the TM of both primers does not differ from each other by more than 5°C. After all, you must choose only one temperature to perform the annealing step. The bigger the difference, the more bias you have towards one primer or the other.
Optimizing denaturation temperature
For example, we know that the denaturation temperature should be as low as reasonable to denature template DNA and thus allowing efficient primer annealing and strand extension1 . At the same time, it should not be too high to prevent loss of enzyme activity, since the half-life of Taq DNA polymerases drastically reduces with increasing temperature2 . Naturally, decades of improvements have resulted in more and more stable PCR reagent formulations – increasing fidelity, specificity and speed performance of enzymes. Thus, the denaturation temperature generally falls between 92°C to 98°C, offering choices based on users’ applications. For example, difficult templates such as GC-rich sequences require a higher denaturation temperature, which not only helps separate the strands, but can also lead to higher specificity in PCR amplification3 .
Optimizing annealing temperature (TA)
Things become more complicated when choosing the right annealing temperature. The closest reference one has is the melting temperatures (TM) of the forward and reverse primers. TM describe the temperature at which there is an equilibrium between the primer bound to DNA and the primer dissociating from DNA, resulting in 50% of the template having bound primer. When choosing primer pairs, it is important that the TM of both primers does not differ from each other by more than 5°C. After all, you must choose only one temperature to perform the annealing step. The bigger the difference, the more bias you have towards one primer or the other.
Read more
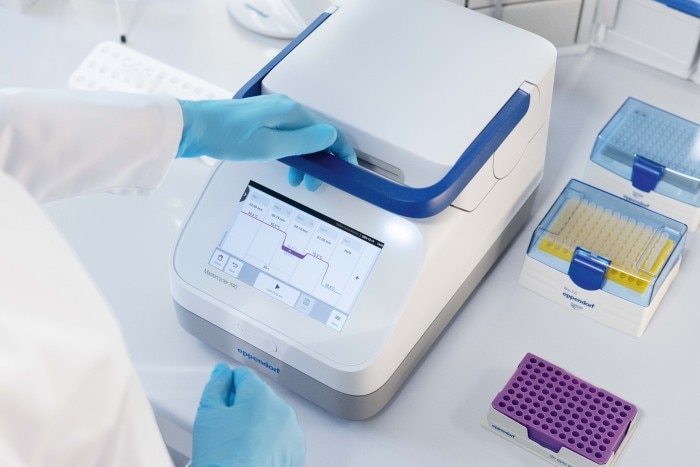

Unfortunately, the TM can change with reagent concentration, pH and salt concentration. Hence, TM remains a theoretical value that can only provide a rough guideline to determine the actual annealing temperature (TA). Using a higher TA results in higher specificity but may affect the yield as it limits binding under more stringent conditions. Thus, finding the most efficient and specific TA is often a trial and error process. This was the main driver behind the innovation of “gradient technology”.
While the extension temperature is typically set at 72 °C for Taq polymerase, optimizing the elongation time in PCR is just as critical for successful amplification. Elongation time determines how long the polymerase has to synthesize the target DNA fragment. A common guideline is ~1 minute per kilobase of target length, but this can vary depending on enzyme speed and template complexity.
Shorter elongation times can significantly reduce total PCR duration, which is advantageous for high-throughput workflows. However, if the target sequence is long or GC-rich, insufficient elongation time may lead to incomplete products. Conversely, longer elongation times improve yield and fidelity but increase cycle time.
Optimizing elongation time in PCR
While the extension temperature is typically set at 72 °C for Taq polymerase, optimizing the elongation time in PCR is just as critical for successful amplification. Elongation time determines how long the polymerase has to synthesize the target DNA fragment. A common guideline is ~1 minute per kilobase of target length, but this can vary depending on enzyme speed and template complexity.
Shorter elongation times can significantly reduce total PCR duration, which is advantageous for high-throughput workflows. However, if the target sequence is long or GC-rich, insufficient elongation time may lead to incomplete products. Conversely, longer elongation times improve yield and fidelity but increase cycle time.
Read more
What is gradient technology and how is it used to optimize PCR conditions?
Gradient PCR is a technology where instead of giving the same temperature throughout the entire thermal block of a thermal cycler , each of the columns or rows in the block is made to have different temperatures. This means that you can either choose to run repeated PCRs testing different temperatures – each about an hour long – to find the right TA. Or, you can choose to use a thermal cycler equipped with gradient function . With it, you can run different temperatures all at once, saving you much time and effort.
Read more
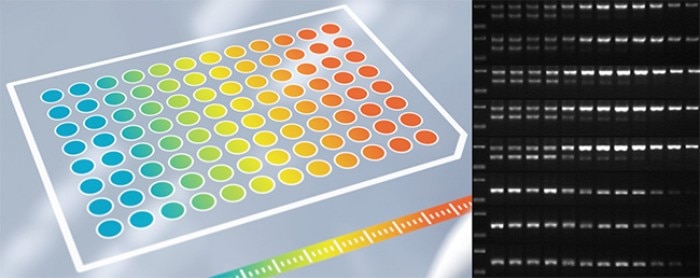

Figure 2: Temperature gradient in PCR, with gel electrophoresis showing how yield and specificity can be affected with varied temperatures.
While gradient PCR primarily focuses on optimizing annealing temperature, it is also an opportunity to review other parameters such as the denaturation temperature. The elongation time can also be optimized in parallel, by running multiple gradient PCRs with varied extension durations.
The analysis step, e.g. gel electrophoresis, is then used to determine the best temperatures and elongation time in PCR that provides the best yield, specificity, and efficiency. Of course, the higher the number of conditions you can run at one go, the more you know about your primer behaviour, making future optimization work easier, resulting in potential savings in time and money.
You can dive deeper into the principles of gradient PCR in our blog and application note .
References:
[1] McPherson, M. & Moller, S. (2006) PCR 2nd edition (https://saddam2014.files.wordpress.com/2014/01/pcr-taylor-francis-2006.pdf)
[2] Innis, M., & Gelfand, D. H. (1999) Optimization of PCRs (http://www.ispybio.com/search/protocols/PCR%20protocol9.pdf)
[3] Gerke, N. & Hellberg, A. (2013) Eppendorf Application Note 289
The analysis step, e.g. gel electrophoresis, is then used to determine the best temperatures and elongation time in PCR that provides the best yield, specificity, and efficiency. Of course, the higher the number of conditions you can run at one go, the more you know about your primer behaviour, making future optimization work easier, resulting in potential savings in time and money.
You can dive deeper into the principles of gradient PCR in our blog and application note .
References:
[1] McPherson, M. & Moller, S. (2006) PCR 2nd edition (https://saddam2014.files.wordpress.com/2014/01/pcr-taylor-francis-2006.pdf)
[2] Innis, M., & Gelfand, D. H. (1999) Optimization of PCRs (http://www.ispybio.com/search/protocols/PCR%20protocol9.pdf)
[3] Gerke, N. & Hellberg, A. (2013) Eppendorf Application Note 289
Read more

